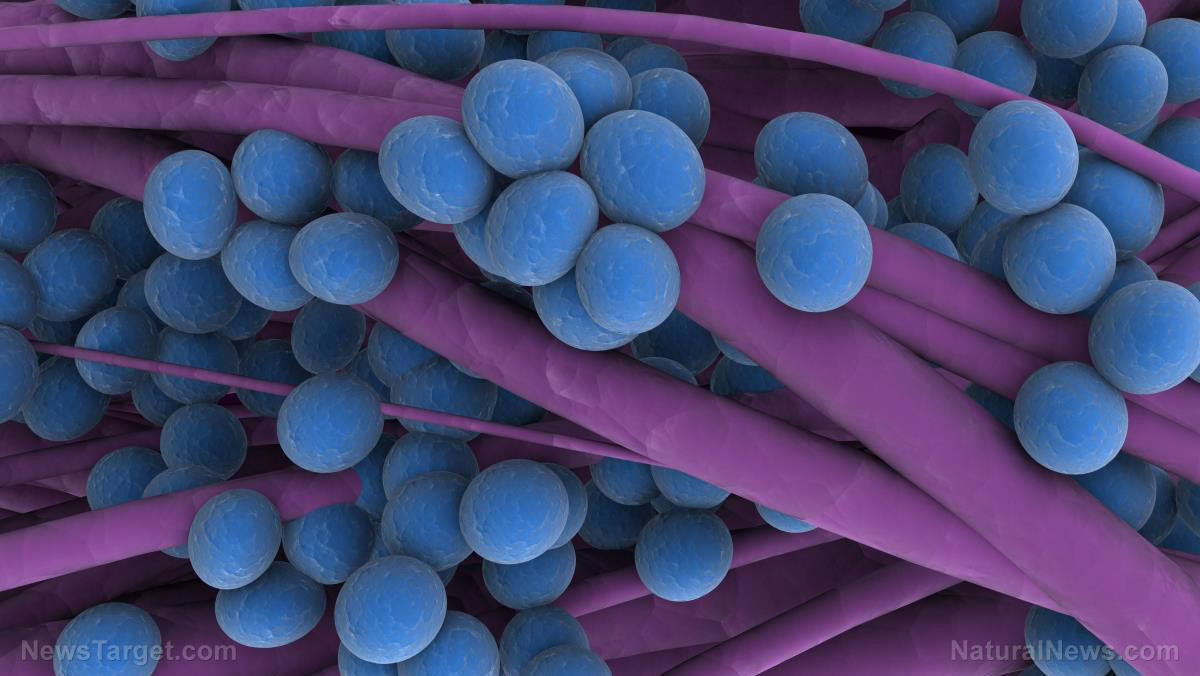
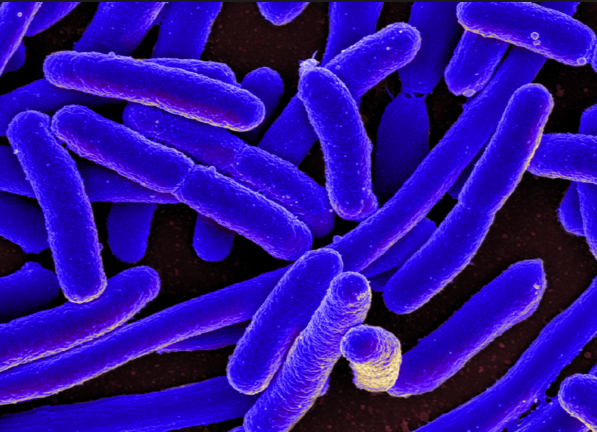
Are we approaching a post-antibiotic future? Chemical medicine reaches “end game” terminal failure
01/30/2017 / By David Gutierrez
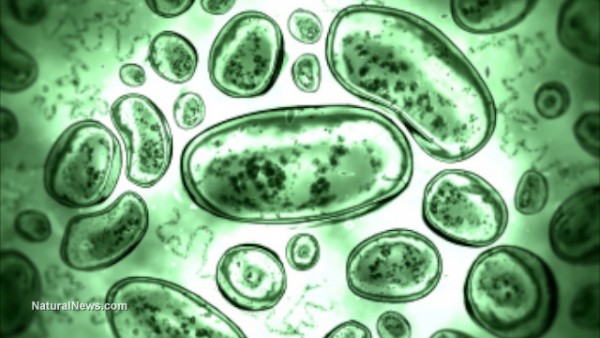
As multi-drug resistance continues to spread throughout bacterial populations, we seem inexorably headed toward a future in which antibiotics cease to be of any use whatsoever, experts have warned.
Antibiotic resistance is a natural product of bacterial evolution. Throughout their history, many bacteria have evolved some degree of resistance to toxic chemicals in their environments, which include the defensive chemicals that other organisms produce against them — the very chemicals that antibiotics are based on. So, when a bacterial population is exposed to antibiotics, all the vulnerable organisms die, leaving only the drug-resistant ones to pass their genes on to the next generation.
But this process has been dramatically accelerated by irresponsible use of antibiotics to treat viral infections or non-dangerous bacterial infections, and particularly by the use of antibiotics as growth-promoters in livestock.
‘Nightmare bacteria’
The future was foreshadowed by a shocking case in August, of a 70-year old woman who was admitted to a Reno hospital suffering septic shock from an infection of Klebsiella pneumoniae. After she died from the infection — which failed to respond to antibiotic treatment — samples of the bacteria were sent to the Centers for Disease Control and Prevention (CDC) for analysis.
Researchers found that the strain of K. pneumoniae was resistant to all 26 antibiotics approved for human use in the United States, including the “drug of last resort,” colistin.
K. pneumoniae is naturally present in the gut, and typically only causes infection in people who are weakened by injury or disease, such as hospital patients. The woman had initially acquired the infection while hospitalized for a broken leg in India. Thus, the “nightmare bacteria,” as the CDC calls it, does not appear to be endemic to the United States — yet.
“The ease of global travel does mean that such cases will increase,” said David Brown, chief scientist at Antibiotic Research UK.
The end of modern medicine
The post-antibiotic future may arrive faster than anyone expected, for one simple reason: All antibiotics are derived from only a handful of separate chemical types. And once bacteria develop resistance to one drug, that resistance rapidly generalizes to every drug in the same class.
Additionally, pharmaceutical companies have largely abandoned research into new antibiotics due to low profit margins.
But, ultimately, even the development of new drugs would only slow the end of antibiotic-based medicine. Experts warn that only widespread changes in antibiotic use, including an end to routine use in animal agriculture, can truly stop the problem.
In 2015, World Health Organization head, Margaret Chan, warned that without these changes, we will enter “a post-antibiotic era, in which common infections will once again kill.”
This era will likely arrive within the lifetimes of many people who are alive today. According to projections by the British government’s Review on Antimicrobial Resistance, by 2050, superbugs will be killing 10 million people per year, more than the deaths from cancer and diabetes combined. That is 10 times the number being killed by these infections today. In some countries, such as Nigeria, it is predicted that superbugs will be responsible for a quarter of all deaths by that same year.
Notably, these numbers count deaths from only six drug-resistant pathogens (though not all of them are bacteria): K. pneumoniae, E. coli, MRSA, HIV, TB and malaria.
A post-antibiotic world will not just mean that people will die from infections that can now be treated. It will mean that many now-routine medical procedures will become impossible — including all surgeries, cancer treatments and organ transplants. The deaths caused by the loss of these medical treatments are also not included in the 10 million per year figure.
In Chan’s words, the loss of effective antibiotics will “mean the end of modern medicine as we know it.” (RELATED: See Medicine.news for breaking new on modern medicine’s failures and achievements.)
Sources for this article include:
Tagged Under: Antibiotics, superbugs